-
PDF
- Split View
-
Views
-
Cite
Cite
Sabine S. Jakob, Frank R. Blattner, A Chloroplast Genealogy of Hordeum (Poaceae): Long-Term Persisting Haplotypes, Incomplete Lineage Sorting, Regional Extinction, and the Consequences for Phylogenetic Inference, Molecular Biology and Evolution, Volume 23, Issue 8, August 2006, Pages 1602–1612, https://doi.org/10.1093/molbev/msl018
Close - Share Icon Share
Abstract
To analyze reasons for inconclusive results of earlier chloroplast phylogenies in the grass genus Hordeum, we established a genealogy of chloroplast haplotypes by sequencing the trnL–trnF region in 875 individuals, covering all 31 species of the genus. Although the outcomes of phenetic and parsimony analyses of 88 haplotypes were ambiguous, a network approach showed that in Hordeum ancient chloroplast types co-occur with their descendants. Moreover, we found up to 18 different chloroplast haplotypes within a single species and up to 6 species sharing single haplotypes. Persisting polymorphisms together with incomplete lineage sorting occurred preferentially in the rapidly speciating New World taxa of the genus, where ancient chloroplast types have survived for at least 4 Myr. Lineages-through-time plots and a high number of missing chloroplast haplotypes indicated far-reaching extinction of chloroplast lineages in Europe and particularly the Mediterranean. Survival of these lineages in East Asia and North America resulted in chloroplast relationships that markedly differed from nuclear estimations of species relationships. Thus, even for the deepest splits in the genus, reaching back more than 9 Myr, no safe phylogenetic inference from chloroplast data is possible in Hordeum. The chloroplast genealogy, however, revealed biogeographic patterns and indicated processes involved in speciation in Hordeum. We conclude that the described phenomena are not restricted to Hordeum and that the knowledge of the chloroplast relationships within a genus is indispensable to prevent misinterpretation of phylogeographic data within single species.
Introduction
Phylogenetic analyses at the interspecific level differ markedly from those within species. Although long-term reproductive isolation is thought to result ultimately in hierarchically structured, nonoverlapping gene pools of different species (phylogeny), intraspecific relationships are characterized by reticulations (tokogeny) and thus are not hierarchical (Hennig 1950). Accordingly, different analytical tools were developed to account for phylogenetic versus tokogenetic relationships. In phylogenetic analyses, branching diagrams, that is, phylogenetic trees, describe the hierarchical patterns of ancestry among different species (Swofford et al. 1996), whereas networks are mostly used to describe tokogenetic patterns (Posada and Crandall 2001). Although problems related to the transition zone between both levels of relationships were long recognized (e.g., Maddison 1997; Avise 2000), algorithms assuming bifurcating data structures are predominantly used to analyze species-level phylogenies.
The barley genus Hordeum comprises 31 species, including diploid (2n = 2x = 14) and polyploid (2n = 4x = 28, 2n = 6x = 42) species or cytotypes (Bothmer et al. 1995). The genus originated about 12 MYA (Blattner 2004) in western Eurasia from where it colonized its extant distribution area in Europe, central Asia, North America, South America, and South Africa (Blattner 2006). The monophyly of the genus is well supported by morphological (Seberg and Frederiksen 2001) and molecular phylogenetic studies (Petersen and Seberg 1997; Blattner 2004), whereas the intrageneric phylogeny is still a matter of dispute. Although the combined analysis of sequences from 3 nuclear loci of all diploid Hordeum species resulted in a well-supported and highly resolved phylogeny (Blattner 2006), studies of the maternally inherited chloroplast genome (Baum and Bailey 1991; Doebley et al. 1992; Provan et al. 1999; Nishikawa et al. 2002) or combined chloroplast and nuclear data sets (Petersen and Seberg 2003) arrived at contradictory phylogenetic hypotheses of species relationships. These inconsistencies occurred even when identical taxon sets were used. Reasons for these far-reaching discrepancies and contradictions in Hordeum were never fully understood.
To analyze the chloroplast relationships in Hordeum, we sampled, wherever available, several individuals per species and sequenced the chloroplast trnL-F region (Taberlet et al. 1991). This region includes the intron of the trnL(UAA) gene and the trnL–trnF(GAA) intergenic spacer. The trnL-F region shows high mutation rates and is often used in phylogenetic and phylogeographic analyses of plants on or below the species level (e.g., Fujii et al. 1997; Bakker et al. 1998; Bleeker et al. 2002; Albach et al. 2004; Koch et al. 2006). To understand the processes resulting in the patterns found in the chloroplast DNA, we compared them with the species' phylogeny derived from nuclear markers (Blattner 2004, 2006). Because phylogenetic analysis algorithms might face problems when data do not represent a tree-like structure (Posada and Crandall 2001), we analyzed the sequences in addition with a statistical parsimony network approach. It reflects genealogical relationships of the chloroplast haplotypes, that is, single mutation steps separate adjacent haplotypes in the network, and older haplotypes are placed at internal branching points whereas younger occur toward tip positions. With this analysis of Hordeum chloroplasts we show that deep coalescence (Hudson 1990) combined with incomplete sorting of ancient lineages can result in incongruence between nuclear and chloroplast phylogenies not only in young and rapidly speciating groups (Mason-Gamer et al. 1995; Comes and Abbott 2001) but also in old lineages reaching deep into the history of a genus. Furthermore, we intend to analyze biogeographical patterns in the chloroplasts to see if the chloroplast data support the biogeographical scenario of the genus derived from nuclear data (Blattner 2006). Because we are interested in the reasons for a rapid radiation of Hordeum taxa in the New World, we also like to see if the chloroplast data can contribute to the understanding of speciation mechanisms in Hordeum.
Materials and Methods
Taxon Sampling
We included 875 individuals obtained from 249 natural populations, 166 lines from germplasm repositories, and 13 herbarium specimens, representing the 31 Hordeum species and nearly all subspecific taxa (table 1). The collection sites of the samples cover the entire distribution area of the genus. For species with cytotypes with different ploidy levels, these were mostly determined by flow cytometric measurement of the genome size of the analyzed individuals (Jakob et al. 2004). The Hordeum accessions, their origins, collectors, and voucher data are listed in table S1 (Supplementary Material online or directly from the authors).
Hordeum Species Studied, Summary of Statistical Parameters, and Chloroplast Haplotype Distribution among Species
Species . | PL . | Ni . | Np . | Distribution . | K . | GDI . | SD . | Haplotypes . |
|---|---|---|---|---|---|---|---|---|
| Hordeum arizonicum Covas | 6x | 2 | 2 | NAm | H | 0 | 0 | 52 |
| Hordeum bogdanii Wilensky | 2x | 3 | 3 | c As | H | 0.67 | 0.31 | 2, 6 |
| Hordeum brachyantherum Nevski | 2x/4x | 54 | 32 | NAm | H | 0.80 | 0.03 | 24, 31, 33, 35, 36, 37, 38, 39, 40, 41 |
| Hordeum brachyantherum Nevski | 6x | 2 | 2 | NAm | HXa | 0 | 0 | 22 |
| Hordeum brevisubulatum (Trin.) Link | 2x/4x/6x | 16 | 12 | c As | H | 0.88 | 0.06 | 3, 4, 5, 7, 8, 9, 10, 11, 14 |
| Hordeum bulbosum L. | 2x/4x | 15 | 10 | sw As, Med | I | 0.73 | 0.09 | 26, 27, 28, 29, 30 |
| Hordeum capense Thunb. | 4x | 1 | 1 | SAf | HXa | — | — | 16 |
| Hordeum chilense Roemer & Schultes | 2x | 21 | 21 | SAm/n | H | 0.18 | 0.10 | 47, 53 |
| Hordeum comosum Presl | 2x | 43 | 16 | SAm/s | H | 0.72 | 0.04 | 46, 47, 56, 57, 60, 65 |
| Hordeum cordobense Bothmer, Jacobsen & Nicora | 2x | 11 | 9 | SAm/n | H | 0.55 | 0.07 | 66, 67 |
| Hordeum depressum (Scribn. & J. G. Sm.) Rydb. | 4x | 3 | 3 | NAm | H | 0.67 | 0.31 | 1, 38 |
| Hordeum erectifolium Bothmer, Jacobsen & Jørgensen | 2x | 1 | 1 | SAm/n | H | — | — | 48 |
| Hordeum euclaston Steud. | 2x | 7 | 7 | SAm/n | H | 0.29 | 0.20 | 55, 56 |
| Hordeum flexuosum Nees ex Steud. | 2x | 3 | 2 | SAm/n | H | 0 | 0 | 46 |
| Hordeum fuegianum Bothmer, Jacobsen & Jørgensen | 4x | 2 | 2 | SAm/s | H | 1 | 0.5 | 81, 82 |
| Hordeum guatemalense Bothmer, Jacobsen & Jørgensen | 4x | 1 | 1 | c America | H | — | — | 38 |
| Hordeum intercedens Nevski | 2x | 3 | 2 | NAm | H | 0 | 0 | 63 |
| Hordeum jubatum L. | 4x | 17 | 14 | NAm | H | 0.46 | 0.13 | 32, 34, 41 |
| Hordeum lechleri (Steud.) Schenck | 6x | 199 | 43 | SAm/s | H | 0.34 | 0.04 | 41, 62, 68, 82, 85 |
| Hordeum marinum Huds. subsp. gussoneanum (Parl.) Thell. | 2x | 34 | 18 | Eu, Med | Xa | 0 | 0 | 22 |
| Hordeum marinum Huds. subsp. gussoneanum (Parl.) Thell. | 4x | 7 | 7 | Eu, Med | Xa | 0 | 0 | 23 |
| Hordeum marinum Huds. subsp. marinum | 2x | 101 | 47 | Eu, Med | Xa | 0.42 | 0.06 | 16, 17, 18, 19, 20, 21 |
| Hordeum murinum L. | 2x | 9 | 8 | Eu, Med | Xu | 0 | 0 | 13 |
| Hordeum murinum L. | 4x/6x | 22 | 20 | Eu, Med | Xu | 0 | 0 | 12 |
| Hordeum muticum J. Presl | 2x | 10 | 10 | SAm/n | H | 0.64 | 0.15 | 1, 43, 44, 45 |
| Hordeum parodii Covas | 6x | 31 | 11 | SAm/s | H | 0.55 | 0.09 | 47, 68, 85, 87 |
| Hordeum patagonicum (Haumann) Covas | 2x | 74 | 38 | SAm/s | H | 0.87 | 0.03 | 46, 54, 57, 59, 62, 64, 68, 70, 71, 72, 73, 76, 77, 78, 79, 83, 85, 88 |
| Hordeum procerum Nevski | 6x | 4 | 4 | SAm/n | H | 0.5 | 0.27 | 1, 42 |
| Hordeum pubiflorum Hook. f. | 2x | 101 | 39 | SAm/s | H | 0.76 | 0.03 | 46, 47, 54, 57, 64, 68, 69, 74, 75, 77, 80, 82, 85, 86 |
| Hordeum pusillum Nutt. | 2x | 3 | 3 | NAm | H | 0 | 0 | 47 |
| Hordeum roshevitzii Bowden | 2x | 2 | 2 | c As | H | 0 | 0 | 5 |
| Hordeum secalinum Schreb. | 4x | 11 | 4 | Eu, Med | HXa | 0 | 0 | 15 |
| Hordeum stenostachys Godr. | 2x | 21 | 16 | SAm/n | H | 0.46 | 0.1 | 49, 50, 51 |
| Hordeum tetraploidum Covas | 4x | 27 | 7 | SAm/s | H | 0.79 | 0.07 | 41, 46, 61, 62, 68, 73, 75, 82, 84 |
| Hordeum vulgare L. | 2x | 14 | 11 | sw As, Med | I | 0 | 0 | 25 |
Species . | PL . | Ni . | Np . | Distribution . | K . | GDI . | SD . | Haplotypes . |
|---|---|---|---|---|---|---|---|---|
| Hordeum arizonicum Covas | 6x | 2 | 2 | NAm | H | 0 | 0 | 52 |
| Hordeum bogdanii Wilensky | 2x | 3 | 3 | c As | H | 0.67 | 0.31 | 2, 6 |
| Hordeum brachyantherum Nevski | 2x/4x | 54 | 32 | NAm | H | 0.80 | 0.03 | 24, 31, 33, 35, 36, 37, 38, 39, 40, 41 |
| Hordeum brachyantherum Nevski | 6x | 2 | 2 | NAm | HXa | 0 | 0 | 22 |
| Hordeum brevisubulatum (Trin.) Link | 2x/4x/6x | 16 | 12 | c As | H | 0.88 | 0.06 | 3, 4, 5, 7, 8, 9, 10, 11, 14 |
| Hordeum bulbosum L. | 2x/4x | 15 | 10 | sw As, Med | I | 0.73 | 0.09 | 26, 27, 28, 29, 30 |
| Hordeum capense Thunb. | 4x | 1 | 1 | SAf | HXa | — | — | 16 |
| Hordeum chilense Roemer & Schultes | 2x | 21 | 21 | SAm/n | H | 0.18 | 0.10 | 47, 53 |
| Hordeum comosum Presl | 2x | 43 | 16 | SAm/s | H | 0.72 | 0.04 | 46, 47, 56, 57, 60, 65 |
| Hordeum cordobense Bothmer, Jacobsen & Nicora | 2x | 11 | 9 | SAm/n | H | 0.55 | 0.07 | 66, 67 |
| Hordeum depressum (Scribn. & J. G. Sm.) Rydb. | 4x | 3 | 3 | NAm | H | 0.67 | 0.31 | 1, 38 |
| Hordeum erectifolium Bothmer, Jacobsen & Jørgensen | 2x | 1 | 1 | SAm/n | H | — | — | 48 |
| Hordeum euclaston Steud. | 2x | 7 | 7 | SAm/n | H | 0.29 | 0.20 | 55, 56 |
| Hordeum flexuosum Nees ex Steud. | 2x | 3 | 2 | SAm/n | H | 0 | 0 | 46 |
| Hordeum fuegianum Bothmer, Jacobsen & Jørgensen | 4x | 2 | 2 | SAm/s | H | 1 | 0.5 | 81, 82 |
| Hordeum guatemalense Bothmer, Jacobsen & Jørgensen | 4x | 1 | 1 | c America | H | — | — | 38 |
| Hordeum intercedens Nevski | 2x | 3 | 2 | NAm | H | 0 | 0 | 63 |
| Hordeum jubatum L. | 4x | 17 | 14 | NAm | H | 0.46 | 0.13 | 32, 34, 41 |
| Hordeum lechleri (Steud.) Schenck | 6x | 199 | 43 | SAm/s | H | 0.34 | 0.04 | 41, 62, 68, 82, 85 |
| Hordeum marinum Huds. subsp. gussoneanum (Parl.) Thell. | 2x | 34 | 18 | Eu, Med | Xa | 0 | 0 | 22 |
| Hordeum marinum Huds. subsp. gussoneanum (Parl.) Thell. | 4x | 7 | 7 | Eu, Med | Xa | 0 | 0 | 23 |
| Hordeum marinum Huds. subsp. marinum | 2x | 101 | 47 | Eu, Med | Xa | 0.42 | 0.06 | 16, 17, 18, 19, 20, 21 |
| Hordeum murinum L. | 2x | 9 | 8 | Eu, Med | Xu | 0 | 0 | 13 |
| Hordeum murinum L. | 4x/6x | 22 | 20 | Eu, Med | Xu | 0 | 0 | 12 |
| Hordeum muticum J. Presl | 2x | 10 | 10 | SAm/n | H | 0.64 | 0.15 | 1, 43, 44, 45 |
| Hordeum parodii Covas | 6x | 31 | 11 | SAm/s | H | 0.55 | 0.09 | 47, 68, 85, 87 |
| Hordeum patagonicum (Haumann) Covas | 2x | 74 | 38 | SAm/s | H | 0.87 | 0.03 | 46, 54, 57, 59, 62, 64, 68, 70, 71, 72, 73, 76, 77, 78, 79, 83, 85, 88 |
| Hordeum procerum Nevski | 6x | 4 | 4 | SAm/n | H | 0.5 | 0.27 | 1, 42 |
| Hordeum pubiflorum Hook. f. | 2x | 101 | 39 | SAm/s | H | 0.76 | 0.03 | 46, 47, 54, 57, 64, 68, 69, 74, 75, 77, 80, 82, 85, 86 |
| Hordeum pusillum Nutt. | 2x | 3 | 3 | NAm | H | 0 | 0 | 47 |
| Hordeum roshevitzii Bowden | 2x | 2 | 2 | c As | H | 0 | 0 | 5 |
| Hordeum secalinum Schreb. | 4x | 11 | 4 | Eu, Med | HXa | 0 | 0 | 15 |
| Hordeum stenostachys Godr. | 2x | 21 | 16 | SAm/n | H | 0.46 | 0.1 | 49, 50, 51 |
| Hordeum tetraploidum Covas | 4x | 27 | 7 | SAm/s | H | 0.79 | 0.07 | 41, 46, 61, 62, 68, 73, 75, 82, 84 |
| Hordeum vulgare L. | 2x | 14 | 11 | sw As, Med | I | 0 | 0 | 25 |
NOTE.—PL, ploidy level; Ni, number of individuals; Np, number of populations/accessions; K, karyotype (genome); GDI, Nei's gene diversity index; SD, standard deviation; As, Asia; Eu, Europe; Med, Mediterranean; NAm, North America; SAm/n, South America/northern group; SAm/s, South America/southern group; SAf, South Africa; c, central; sw, southwest.
Hordeum Species Studied, Summary of Statistical Parameters, and Chloroplast Haplotype Distribution among Species
Species . | PL . | Ni . | Np . | Distribution . | K . | GDI . | SD . | Haplotypes . |
|---|---|---|---|---|---|---|---|---|
| Hordeum arizonicum Covas | 6x | 2 | 2 | NAm | H | 0 | 0 | 52 |
| Hordeum bogdanii Wilensky | 2x | 3 | 3 | c As | H | 0.67 | 0.31 | 2, 6 |
| Hordeum brachyantherum Nevski | 2x/4x | 54 | 32 | NAm | H | 0.80 | 0.03 | 24, 31, 33, 35, 36, 37, 38, 39, 40, 41 |
| Hordeum brachyantherum Nevski | 6x | 2 | 2 | NAm | HXa | 0 | 0 | 22 |
| Hordeum brevisubulatum (Trin.) Link | 2x/4x/6x | 16 | 12 | c As | H | 0.88 | 0.06 | 3, 4, 5, 7, 8, 9, 10, 11, 14 |
| Hordeum bulbosum L. | 2x/4x | 15 | 10 | sw As, Med | I | 0.73 | 0.09 | 26, 27, 28, 29, 30 |
| Hordeum capense Thunb. | 4x | 1 | 1 | SAf | HXa | — | — | 16 |
| Hordeum chilense Roemer & Schultes | 2x | 21 | 21 | SAm/n | H | 0.18 | 0.10 | 47, 53 |
| Hordeum comosum Presl | 2x | 43 | 16 | SAm/s | H | 0.72 | 0.04 | 46, 47, 56, 57, 60, 65 |
| Hordeum cordobense Bothmer, Jacobsen & Nicora | 2x | 11 | 9 | SAm/n | H | 0.55 | 0.07 | 66, 67 |
| Hordeum depressum (Scribn. & J. G. Sm.) Rydb. | 4x | 3 | 3 | NAm | H | 0.67 | 0.31 | 1, 38 |
| Hordeum erectifolium Bothmer, Jacobsen & Jørgensen | 2x | 1 | 1 | SAm/n | H | — | — | 48 |
| Hordeum euclaston Steud. | 2x | 7 | 7 | SAm/n | H | 0.29 | 0.20 | 55, 56 |
| Hordeum flexuosum Nees ex Steud. | 2x | 3 | 2 | SAm/n | H | 0 | 0 | 46 |
| Hordeum fuegianum Bothmer, Jacobsen & Jørgensen | 4x | 2 | 2 | SAm/s | H | 1 | 0.5 | 81, 82 |
| Hordeum guatemalense Bothmer, Jacobsen & Jørgensen | 4x | 1 | 1 | c America | H | — | — | 38 |
| Hordeum intercedens Nevski | 2x | 3 | 2 | NAm | H | 0 | 0 | 63 |
| Hordeum jubatum L. | 4x | 17 | 14 | NAm | H | 0.46 | 0.13 | 32, 34, 41 |
| Hordeum lechleri (Steud.) Schenck | 6x | 199 | 43 | SAm/s | H | 0.34 | 0.04 | 41, 62, 68, 82, 85 |
| Hordeum marinum Huds. subsp. gussoneanum (Parl.) Thell. | 2x | 34 | 18 | Eu, Med | Xa | 0 | 0 | 22 |
| Hordeum marinum Huds. subsp. gussoneanum (Parl.) Thell. | 4x | 7 | 7 | Eu, Med | Xa | 0 | 0 | 23 |
| Hordeum marinum Huds. subsp. marinum | 2x | 101 | 47 | Eu, Med | Xa | 0.42 | 0.06 | 16, 17, 18, 19, 20, 21 |
| Hordeum murinum L. | 2x | 9 | 8 | Eu, Med | Xu | 0 | 0 | 13 |
| Hordeum murinum L. | 4x/6x | 22 | 20 | Eu, Med | Xu | 0 | 0 | 12 |
| Hordeum muticum J. Presl | 2x | 10 | 10 | SAm/n | H | 0.64 | 0.15 | 1, 43, 44, 45 |
| Hordeum parodii Covas | 6x | 31 | 11 | SAm/s | H | 0.55 | 0.09 | 47, 68, 85, 87 |
| Hordeum patagonicum (Haumann) Covas | 2x | 74 | 38 | SAm/s | H | 0.87 | 0.03 | 46, 54, 57, 59, 62, 64, 68, 70, 71, 72, 73, 76, 77, 78, 79, 83, 85, 88 |
| Hordeum procerum Nevski | 6x | 4 | 4 | SAm/n | H | 0.5 | 0.27 | 1, 42 |
| Hordeum pubiflorum Hook. f. | 2x | 101 | 39 | SAm/s | H | 0.76 | 0.03 | 46, 47, 54, 57, 64, 68, 69, 74, 75, 77, 80, 82, 85, 86 |
| Hordeum pusillum Nutt. | 2x | 3 | 3 | NAm | H | 0 | 0 | 47 |
| Hordeum roshevitzii Bowden | 2x | 2 | 2 | c As | H | 0 | 0 | 5 |
| Hordeum secalinum Schreb. | 4x | 11 | 4 | Eu, Med | HXa | 0 | 0 | 15 |
| Hordeum stenostachys Godr. | 2x | 21 | 16 | SAm/n | H | 0.46 | 0.1 | 49, 50, 51 |
| Hordeum tetraploidum Covas | 4x | 27 | 7 | SAm/s | H | 0.79 | 0.07 | 41, 46, 61, 62, 68, 73, 75, 82, 84 |
| Hordeum vulgare L. | 2x | 14 | 11 | sw As, Med | I | 0 | 0 | 25 |
Species . | PL . | Ni . | Np . | Distribution . | K . | GDI . | SD . | Haplotypes . |
|---|---|---|---|---|---|---|---|---|
| Hordeum arizonicum Covas | 6x | 2 | 2 | NAm | H | 0 | 0 | 52 |
| Hordeum bogdanii Wilensky | 2x | 3 | 3 | c As | H | 0.67 | 0.31 | 2, 6 |
| Hordeum brachyantherum Nevski | 2x/4x | 54 | 32 | NAm | H | 0.80 | 0.03 | 24, 31, 33, 35, 36, 37, 38, 39, 40, 41 |
| Hordeum brachyantherum Nevski | 6x | 2 | 2 | NAm | HXa | 0 | 0 | 22 |
| Hordeum brevisubulatum (Trin.) Link | 2x/4x/6x | 16 | 12 | c As | H | 0.88 | 0.06 | 3, 4, 5, 7, 8, 9, 10, 11, 14 |
| Hordeum bulbosum L. | 2x/4x | 15 | 10 | sw As, Med | I | 0.73 | 0.09 | 26, 27, 28, 29, 30 |
| Hordeum capense Thunb. | 4x | 1 | 1 | SAf | HXa | — | — | 16 |
| Hordeum chilense Roemer & Schultes | 2x | 21 | 21 | SAm/n | H | 0.18 | 0.10 | 47, 53 |
| Hordeum comosum Presl | 2x | 43 | 16 | SAm/s | H | 0.72 | 0.04 | 46, 47, 56, 57, 60, 65 |
| Hordeum cordobense Bothmer, Jacobsen & Nicora | 2x | 11 | 9 | SAm/n | H | 0.55 | 0.07 | 66, 67 |
| Hordeum depressum (Scribn. & J. G. Sm.) Rydb. | 4x | 3 | 3 | NAm | H | 0.67 | 0.31 | 1, 38 |
| Hordeum erectifolium Bothmer, Jacobsen & Jørgensen | 2x | 1 | 1 | SAm/n | H | — | — | 48 |
| Hordeum euclaston Steud. | 2x | 7 | 7 | SAm/n | H | 0.29 | 0.20 | 55, 56 |
| Hordeum flexuosum Nees ex Steud. | 2x | 3 | 2 | SAm/n | H | 0 | 0 | 46 |
| Hordeum fuegianum Bothmer, Jacobsen & Jørgensen | 4x | 2 | 2 | SAm/s | H | 1 | 0.5 | 81, 82 |
| Hordeum guatemalense Bothmer, Jacobsen & Jørgensen | 4x | 1 | 1 | c America | H | — | — | 38 |
| Hordeum intercedens Nevski | 2x | 3 | 2 | NAm | H | 0 | 0 | 63 |
| Hordeum jubatum L. | 4x | 17 | 14 | NAm | H | 0.46 | 0.13 | 32, 34, 41 |
| Hordeum lechleri (Steud.) Schenck | 6x | 199 | 43 | SAm/s | H | 0.34 | 0.04 | 41, 62, 68, 82, 85 |
| Hordeum marinum Huds. subsp. gussoneanum (Parl.) Thell. | 2x | 34 | 18 | Eu, Med | Xa | 0 | 0 | 22 |
| Hordeum marinum Huds. subsp. gussoneanum (Parl.) Thell. | 4x | 7 | 7 | Eu, Med | Xa | 0 | 0 | 23 |
| Hordeum marinum Huds. subsp. marinum | 2x | 101 | 47 | Eu, Med | Xa | 0.42 | 0.06 | 16, 17, 18, 19, 20, 21 |
| Hordeum murinum L. | 2x | 9 | 8 | Eu, Med | Xu | 0 | 0 | 13 |
| Hordeum murinum L. | 4x/6x | 22 | 20 | Eu, Med | Xu | 0 | 0 | 12 |
| Hordeum muticum J. Presl | 2x | 10 | 10 | SAm/n | H | 0.64 | 0.15 | 1, 43, 44, 45 |
| Hordeum parodii Covas | 6x | 31 | 11 | SAm/s | H | 0.55 | 0.09 | 47, 68, 85, 87 |
| Hordeum patagonicum (Haumann) Covas | 2x | 74 | 38 | SAm/s | H | 0.87 | 0.03 | 46, 54, 57, 59, 62, 64, 68, 70, 71, 72, 73, 76, 77, 78, 79, 83, 85, 88 |
| Hordeum procerum Nevski | 6x | 4 | 4 | SAm/n | H | 0.5 | 0.27 | 1, 42 |
| Hordeum pubiflorum Hook. f. | 2x | 101 | 39 | SAm/s | H | 0.76 | 0.03 | 46, 47, 54, 57, 64, 68, 69, 74, 75, 77, 80, 82, 85, 86 |
| Hordeum pusillum Nutt. | 2x | 3 | 3 | NAm | H | 0 | 0 | 47 |
| Hordeum roshevitzii Bowden | 2x | 2 | 2 | c As | H | 0 | 0 | 5 |
| Hordeum secalinum Schreb. | 4x | 11 | 4 | Eu, Med | HXa | 0 | 0 | 15 |
| Hordeum stenostachys Godr. | 2x | 21 | 16 | SAm/n | H | 0.46 | 0.1 | 49, 50, 51 |
| Hordeum tetraploidum Covas | 4x | 27 | 7 | SAm/s | H | 0.79 | 0.07 | 41, 46, 61, 62, 68, 73, 75, 82, 84 |
| Hordeum vulgare L. | 2x | 14 | 11 | sw As, Med | I | 0 | 0 | 25 |
NOTE.—PL, ploidy level; Ni, number of individuals; Np, number of populations/accessions; K, karyotype (genome); GDI, Nei's gene diversity index; SD, standard deviation; As, Asia; Eu, Europe; Med, Mediterranean; NAm, North America; SAm/n, South America/northern group; SAm/s, South America/southern group; SAf, South Africa; c, central; sw, southwest.
Molecular Methods
Genomic DNA was extracted from about 50 mg fresh or 10 mg silica-dried leaf material using the DNeasy Plant Kit (Qiagen, Hilden, Germany) according to the instructions of the manufacturer. To analyze the chloroplast trnL–trnF region of Hordeum, polymerase chain reaction (PCR) primers were designed to match the rps4 (5′-CGATAACGGGACATGAAG-3′) and the trnF(GAA) gene (5′-ATTTGAACTGGTGACACGAG-3′) of Hordeum. PCR was performed with a GeneAmp 9700 PCR System (PE Biosystems, Foster City, CA) with the following cycling program: 95 °C for 3 min; 10 cycles of 95 °C for 30 s, 57 °C for 35 s, 68 °C for 2.3 min; and 30 cycles of 95 °C for 30 s, 54 °C for 35 s, and 68 °C for 2.3 min, followed by a posttreatment at 68 °C for 15 min. The reaction was carried out with 1.5 U Taq DNA polymerase (Qiagen) in the supplied reaction buffer, 0.2 mM of each deoxynucleoside triphosphate, 25 pmol of each primer, and about 20 ng of total DNA in a final reaction volume of 50 μl. To weaken DNA secondary structures during PCR, Q-Solution (Qiagen) was added to a final concentration of 20%.
The amplicons of about 3500 bp were purified using Nucleofast PCR Plates (Macherey-Nagel, Düren, Germany) and resuspended in 35 μl TE buffer. For cycle sequencing, we used a nested forward primer (5′-GGAAATGGGGATATGGCG-3′) that anneals in exon 1 of the trnL(UAA) gene, whereas the reverse primer was the trnF primer used for PCR amplification. This resulted in sequences of the trnL gene and the trnL–trnF intergenic spacer. Sequencing about 900–1000 bp out of the 3500-bp amplicon allows us to easily increase sequence length for crucial taxa without the need of new DNA amplification and purification. PCR products were directly sequenced on a MegaBACE 1000 automatic DNA sequencer using the respective dye-terminator sequencing technology (Amersham Biosciences, Freiburg, Germany) with the addition of Q-Solution (Qiagen) to a final concentration of 10%. Forward and reverse sequences of each sample were manually edited and combined into single consensus sequences.
Data Analyses
Aligning of the sequences could be done manually by introducing gaps of 1–18 bp length (in the tetraploid form of Hordeum marinum subsp. gussoneanum, one 73 bp insertion was required) at 21 positions. Length variation at 3 mononucleotide repeats (2 T/A and 1 C/G) was excluded from the analyses due to uncertain homology of the sequence positions. Identical sequences were grouped into haplotypes. The appropriate model of sequence evolution was inferred in Modeltest 3.06 with the Akaike information criterion (Posada and Crandall 1998). We used 2 different analytical approaches. Phylogenetic trees were calculated in PAUP* 4.0b10 (Swofford 2002) using Neighbor-Joining (NJ) analysis of pairwise genetic distances and maximum parsimony analysis with the heuristic search algorithm and the 2-step search strategy given by Blattner (2004), restricting the number of equally most parsimonious trees retained to 80 000. Bootstrap support of branches was calculated for the NJ analysis with 1000 data resamples. In the phylogenetic analyses, we included one sequence from each species possessing a specific chloroplast haplotype, that is, 125 sequences. TCS 1.13 (Clement et al. 2000) was used for a statistical parsimony network approach. Each insertion/deletion (indel) was considered as a single mutation event, and all indels were therefore coded as single positions in the final alignment. In cases where it was not possible to code the indel variation by a single alignment position (overlapping indels or point mutations in insertions), the network was calculated with the necessary number of alignment positions to represent all variation at these sites, and mutation steps were afterward adjusted manually. Here, each haplotype was included only once.
To analyze species diversification rates, an ultrametric tree of diploid Hordeum species (Blattner 2006) was adopted to obtain lineages-through-time plots (Hey 1992; Baldwin and Sanderson 1998; Barraclough and Nee 2001) for the entire genus and the Old and New World taxa separately. This tree was derived from a penalized likelihood approach on the combined sequences from 3 nuclear loci (disrupted meiotic cDNA 1, transcription elongation factor γ, and nrDNA internal transcribed spacer) with the truncated Newton algorithm in r8s (Sanderson 2002), fixing the crown age of Hordeum at 12 Myr (Blattner 2006). The age was obtained from the divergence time of the wheat and barley lineages of 13 Myr (Gaut 2002). To allow comparisons with other taxa, speciation rates per million years were calculated under a Yule process using the Kendall/Moran estimator (Baldwin and Sanderson 1998). To account for errors in the divergence times themselves, the speciation rates were recalculated for the minimum and maximum branch lengths (derived from a parametric bootstrap approach) given by Blattner (2006).
To have a measure for the probability that 2 randomly chosen haplotypes were different within species, Nei's (1987) gene diversity was calculated for all species in ARLEQUIN 2.0 (Schneider et al. 2000).
Results
Haplotype Sequences
The sequenced trnL–trnF region of the Hordeum individuals was in a range between 899 and 989 bp, resulting in an alignment of 1079 bp length. Excluding the length variation at 3 mononucleotide repeats resulted in an alignment length of 1053 bp (used in the phylogenetic analyses). Treating, in addition, every indel as a single mutational event shortened the alignment to 877 bp with 83 variable positions (used in the network approach). Eighty-eight chloroplast haplotypes were identified (EMBL nucleotide database accession numbers AJ969264–AJ969333, AJ969334–AJ969340, AJ969342–AJ969344, AJ969346–AJ969352, AJ969354–AJ969403, and AM231034–AM231036). Seventy of the 88 haplotypes were exclusive for single species, whereas 18 haplotypes were shared among up to 6 species. The haplotype distribution among the Hordeum species and Nei's gene diversity indices for each species are given in table 1.
Phylogenetic Analyses
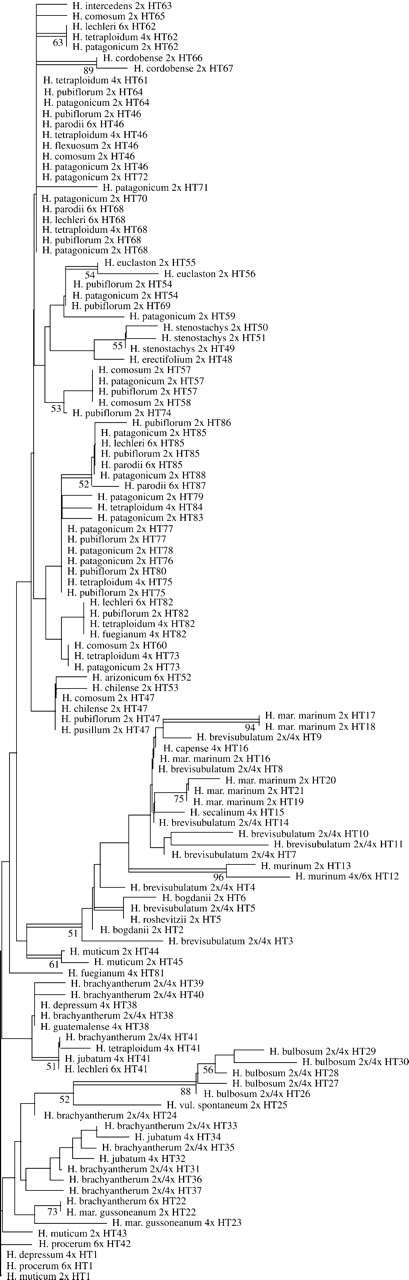
NJ analysis of F81uf + I + Γ genetic distances of the chloroplast haplotype sequences resulted in the tree given in figure 1. A high number of zero-length branches occurred and were often combined with polytomies at these nodes. Maximum parsimony analysis resulted in more than 100,000 most parsimonious trees and could not run to completion. The strict consensus tree of a 2-step search restricted to 80,000 trees (length 107 steps, consistency index = 0.717, retention index = 0.891) was mainly unresolved. Groups retained in this tree are also indicated in figure 1.
Unrooted NJ tree of trnL–trnF sequences of 88 chloroplast haplotypes derived from 875 Hordeum individuals covering all species of the genus. The numbers of the haplotypes and the species' ploidy levels are given. Double bars depict groups that were retained in the strict consensus tree of a parsimony analysis, whereas numbers along the branches denote bootstrap values in the NJ analysis.
Statistical Parsimony Network
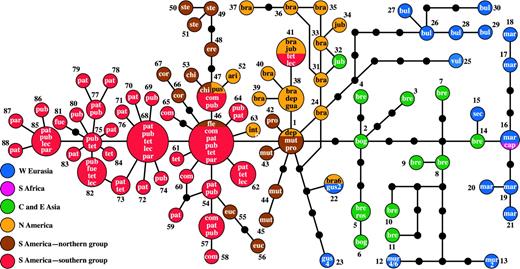
TCS calculated a 95% parsimony connection limit of 12 steps and resulted in a haplotype network of the entire genus Hordeum (fig. 2). Forty-seven haplotypes inferred by TCS were not found in the analyzed individuals and occur as missing intermediates in the network. Five positions in the TCS network revealed closed loops that could not unambiguously be resolved. These loops were each caused by single homoplastic alignment positions.
TCS network of 88 Hordeum chloroplast haplotypes. Circle size represents the number of species sharing specific chloroplast types. Taxon names were abbreviated by the first 3 letters of the (sub)species epithet. The numbers are arbitrary haplotype numbers. Black dots depict missing intermediate haplotypes that were not found in the analyzed individuals.
The haplotype network reveals a clear geographical structure with chloroplast lineages distributed mainly in the Old World or the New World. All lineages from different distribution areas converge at the central haplotype 1 (HT1). A pronounced difference between both groups is the occurrence of 38 missing intermediates in the Old World part of the network as opposed to 9 in the New World part. In the New World, the major division is between North American and South American haplotypes, the latter subdivided into a northern (Chile, central Argentina, Uruguay, and northern Andes) and a southern group (southern Patagonia and Andes), with one chloroplast lineage (HT ≥ 68) exclusive to this southern group. Geographical outliers are the chloroplast haplotype of South African Hordeum capense that is shared with European Hordeum marinum subsp. marinum (HT16), the Siberian sample of Hordeum jubatum (HT32) that falls in a North American Hordeum brachyantherum clade, the occurrence of the otherwise North American HT41 in 2 South American polyploids (Hordeum tetraploidum, Hordeum lechleri), and the South American HT47, 53, and 63 in North American taxa. Also HT22 falls in this pattern as it is shared between a Eurasian diploid (H. marinum subsp. gussoneanum) and its North American allohexaploid descendant (H. brachyantherum 6x). As this allopolyploid originated only in historical times, after the introduction of the diploid into California by humans (Bothmer et al. 1995; Blattner 2004), we will not further discuss this pattern.
Diversification Rates
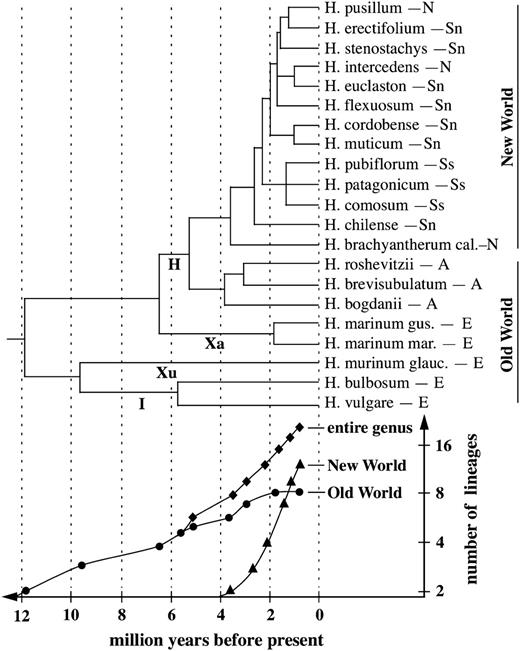
The calculation of speciation rates in diploid Hordeum species assuming a Yule process and an age of the genus of 12 Myr (Blattner 2004, 2006) resulted in a diversification rate of 0.23 ± 0.05 species per million years. However, the lineages-through-time plots (fig. 3) showed that the net diversification rate of the genus during the last 3 Myr was mainly influenced by speciation events in the New World. Accordingly, independent calculations for the Old and New World groups of Hordeum resulted in diversification rates of 0.11 ± 0.03 and 0.42 ± 0.11 species/Myr, respectively.
Phylogenetic tree of the diploid Hordeum species derived from sequences of 3 nuclear loci (Blattner 2006). This tree was adopted for a graphical representation of the diversification rates in Hordeum in a lineages-through-time plot. Letters below the branches depict the Triticeae genomes occurring in Hordeum. The geographical distributions of the species are coded as follows: E = western Eurasia, A = central and East Asia, N = North America, Sn = South America/northern group, and Ss = South America/southern group.
Discussion
Comparison of Analytical Approaches
To analyze 88 haplotype sequences of the chloroplast trnL–trnF region of 875 individuals covering all Hordeum species, we used phylogenetic (i.e., tree based) and genealogical (network) approaches. Maximum parsimony analysis resulted in a poorly resolved tree, and the groups found in the phenetic analysis (NJ) received low bootstrap support (fig. 1), although a certain number of variable positions occur in the trnL-F region. This result is not different from earlier chloroplast analyses in Hordeum (Doebley et al. 1992; Nishikawa et al. 2002; Petersen and Seberg 2003), all of which were characterized by low branch support and nearly no resolution in the New World taxa. Only Provan et al. (1999) obtained a well-resolved NJ tree by combining length differences at 7 chloroplast mononucleotide repeat loci into haplotypes. Unfortunately, no branch support values were reported, and no parsimony analysis was done, which makes comparison with the other studies impossible. Furthermore, all studies arrived at contradictory phylogenetic hypotheses for groups resolved in the trees. Although these studies were relatively complete with respect to the diploid Hordeum species, they included mostly just one individual per taxon or cytotype. In our analysis, the inclusion of several individuals from different origins for most Hordeum species revealed up to 18 chloroplast haplotypes in a single species, and single haplotypes shared among up to 6 species (fig. 2). Chloroplasts were not only shared between (allo)polyploids and their maternal progenitors but also among diploid species. Furthermore, haplotypes from single species mostly occur at different positions in the phylogenetic tree (fig. 1). This combination should result in nearly arbitrary species relationships in phylogenetic analyses, when only one or few individuals are included (Wendel and Doyle 1998), and explains contradictory results in earlier Hordeum phylogenies. As we also found chloroplast mononucleotide repeats to be highly variable and homoplastic for interspecific comparisons (see below), we must conclude that the results of all published chloroplast studies of Hordeum (Baum and Bailey 1991; Doebley et al. 1992; Provan et al. 1999; Nishikawa et al. 2002; Petersen and Seberg 2003) might not be wrong but do not reflect species phylogenies.
Another characteristic of our NJ tree is the occurrence of a high number of zero-length branches in all Hordeum clades, mostly associated with multifurcations. This indicates the persistence of ancestral haplotypes together with their descendants and also that multiple derived lineages evolved from single ancestral chloroplast types (Posada and Crandall 2001). Tree-based analysis methods cannot represent these multifurcating relationships and the coexistence of ancestors with their derivatives, whereas network approaches used in population analyses were designed to deal with such situations. The analysis of the Hordeum chloroplast relationships with TCS resulted in a network where roughly one-third of the haplotypes observed were placed as interiors, partly at multifurcating positions (fig. 2). Five closed loops appeared in the network, which could not be resolved unambiguously. These were caused by single homoplastic sequence positions. Also, we omitted length variation at 3 mononucleotide repeats from the analysis due to uncertain sequence homology. The inclusion of these regions resulted in a large number of closed loops, indicating homoplasious character states (data not shown).
Predictions from coalescent theory assume that older alleles should prevail in populations and be characterized by a higher number of descending lineages and a geographically wider distribution (Crandall and Templeton 1993; Posada and Crandall 2001). In interspecific phylogenies, however, these parameters depend much more on sample design and differences in the history of the taxa. For instance, HT1 may be the oldest extant chloroplast type in our analysis as it connects Hordeum lineages from all continents (fig. 2) and also the major phylogenetic groups as defined by nuclear markers (Blattner 2006) of Hordeum (fig. 3). Its frequency is, however, relatively low in our study. This might partly depend on our sparse sampling in the South American north group and in central Asia. But also in North America it seems to be rare as we found it in allopolyploid Hordeum depressum (4x) but up to now not in the far more extensive sample of its maternal parent H. brachyantherum (Nishikawa et al. 2002). In contrast, in Argentinean H. lechleri (6x) HT41 is the most frequent, occurring invariably in nearly all individuals apart from some in southern most Patagonia. Unlike HT1, this haplotype is in a tip position in our network but is extremely widespread. Thus, we conclude that although we regard genealogical methods to be superior for the analysis of chloroplast lineages in Hordeum, we have to be cautious with coalescence predictions at the interspecific level as differences in life-history traits and generally the fate of the species influence haplotype frequencies.
Mechanisms Resulting in Extant Chloroplast Distribution in Hordeum Species
The chloroplast network of Hordeum is partitioned into New and Old World groups. Shared haplotypes dominate the New World part of the network, whereas in the Old World haplotypes are mostly species specific. The nuclear phylogeny of diploid Hordeum (fig. 3) suggests that the European and Southwest Asian species groups are relatively old lineages whereas the H-genome taxa are younger and underwent a rapid radiation during the last 4 Myr, particularly in South America. Accordingly, net diversification rates are comparatively high (Baldwin and Sanderson 1998) in the New World with 0.42 ± 0.11 species/Myr, especially when taking into account that 40% of the New World species are polyploids and thus not included in the calculation. Diversification was much lower in the Old World with 0.11 ± 0.03 species/Myr. The lineages-through-time plot derived from the nuclear phylogeny of the diploid Hordeum species (fig. 3) revealed a slowdown of net speciation rates during the last 2 Myr for the Old World taxa instead of the expected increase toward the present (Barraclough and Nee 2001; Zhang et al. 2004). This can be due to either decreasing speciation or increasing extinction rates. The high number of missing intermediates in the Old World indicates that far-reaching extinction of Hordeum populations must have taken place in Western Europe, the Mediterranean, and Southwest Asia, simultaneously resulting in the loss of chloroplast haplotypes. The lineages-through-time plot suggests that this happened most probably during the Pleistocene. This extinction scenario fits well to the known impact of the ice age climatic changes on the floristic composition of Europe (Comes and Kadereit 1998). Pleistocene extinctions resulted in shallow coalescence and accordingly mainly species-specific chloroplast lineages in the Old World. In central Asia, this sorting process is either not finished or some hybridization is going on as we found one haplotype (HT5) shared between Hordeum brevisubulatum and Hordeum roshevitzii. Among the Asian species, haplotype diversity is particularly high in the geographically widespread H. brevisubulatum polyploid complex (2x, 4x, and 6x). This can be a result of a pronounced substructuring of the gene pool as cytotypes at different ploidy levels and remote populations are genetically nearly isolated.
Subspecies of H. marinum occupy highly different positions in the chloroplast network, although they are closely related according to nuclear markers (Komatsuda et al. 1999, 2001; Petersen and Seberg 2003; Blattner 2004, 2006). As lateral chloroplast transfer is highly unlikely due to biogeographical reasons and strong homoploid crossing barriers between H. marinum and other Hordeum species, lineage sorting was proposed (Petersen and Seberg 2003) to explain the occurrence of the so-called New World chloroplasts in H. marinum subsp. gussoneanum. From our data, we would further conclude that these chloroplast types (HT22 and 23; fig. 2) are remainders of an ancient stock of Eurasian chloroplast lineages that, apart from the chloroplasts of the H. marinum subsp. gussoneanum, became extinct in the Old World but survived in North America. A similar process, that is, the extinction of ancient chloroplast haplotypes in the other western Eurasian Hordeum taxa, can explain their peripheral position in the network. This, together with the survival of these haplotypes in East Asian and the New World taxa, also explains the inconsistencies between the tree topologies from nuclear and chloroplast data in this group.
Incongruence between phylogenetic data sets derived from the different genomes in plants was mostly explained by cryptic hybridization and introgression (Rieseberg and Soltis 1991; Rieseberg et al. 1996). However, evidence is accumulating that incomplete or differential lineage sorting, that is, the persistence of ancestral polymorphisms through speciation events, also contributes to phylogenetic incongruence (Mason-Gamer et al. 1995; Wendel and Doyle 1998; Comes and Abbott 2001; Linder and Rieseberg 2004). Young species groups in particular should be affected by incomplete lineage sorting while its influence should decrease with increasing time due to the gradual loss of polymorphisms and fixation of lineage-specific alleles (Maddison 1997; Wendel and Doyle 1998). As hybridization also occurs mostly between young and reproductively not completely isolated species, it is often impossible to distinguish the 2 processes in phylogenetic analyses (Wendel and Doyle 1998; Comes and Abbott 2001). The New World group of Hordeum is characterized by many haplotypes shared among species and high chloroplast diversity within species. However, in this rapidly radiating group we have found nearly no indications for ongoing gene flow among diploid species, although the species are relatively young. In the thoroughly sampled Patagonian populations, chloroplast types were mostly distinct in different species where two or more taxa occurred in mixed stands (table 2). In 13 populations with sympatrically occurring diploids, we found only 4 where haplotypes were shared among diploids, and only one of these was a haplotype in a relatively derived position in the network (HT85). Thus, as mostly interior haplotypes, if at all, are shared among diploid species and shared chloroplast types also occur between species without overlapping distribution areas, persisting ancient polymorphisms seem to be the best explanation for the extant chloroplast patterns in diploid Hordeum. Although we cannot completely exclude that hybridization is contributing to the observed haplotype pattern, our data show that only a small amount of haplotypes might have crossed species borders via recent introgression. In contrast to the diploid species, there is some evidence that gene flow occurs between diploid and polyploid species. In mixed populations of southern Patagonia, we found some polyploid individuals that had the chloroplast type of sympatrically occurring diploid Hordeum individuals whereas the majority of the population had a distinct chloroplast type (table 2). We have, however, no indications for chloroplast transfer from polyploids to diploids.
South American Collection Sites with Sympatric Occurrence of Two or More Hordeum Species and Chloroplast Haplotype Distribution among the Taxa
Collection Site . | Species (N) . | Haplotype . | Shared Haplotype . |
|---|---|---|---|
| JB013 | Hordeum pubiflorum (5) | 5 × HT68 | |
| Hordeum patagonicum (7) | 2 × HT54, 1 × HT57, 4 × HT68 | HT68 | |
| Hordeum comosum (3) | 3 × HT46 | ||
| JB014 | Hordeum lechleri (1) | 1 × HT41 | |
| Hordeum parodii (3) | 3 × HT68 | HT57 | |
| Hordeum patagonicum (3) | 3 × HT57 | ||
| Hordeum comosum (3) | 3 × HT57 | ||
| JB015 | Hordeum lechleri (4) | 4 × HT41 | |
| Hordeum pubiflorum (2) | 1 × HT68, 1 × HT85 | HT68 | |
| Hordeum patagonicum (3) | 3 × HT85 | HT85 | |
| Hordeum comosum (3) | 3 × HT57 | ||
| Hordeum parodii (2) | 2 × HT68 | ||
| JB016 | Hordeum lechleri (3) | 3 × HT41 | |
| Hordeum pubiflorum (2) | 2 × HT68 | ||
| JB017 | Hordeum lechleri (4) | 4 × HT41 | |
| Hordeum pubiflorum (6) | 6 × HT68 | ||
| JB019 | Hordeum parodii (5) | 5 × HT46 | |
| Hordeum lechleri (5) | 5 × HT41 | ||
| JB022 | Hordeum pubiflorum (4) | 4 × HT68 | |
| Hordeum comosum (8) | 8 × HT46 | ||
| Hordeum lechleri (3) | 3 × HT41 | ||
| JB025 | Hordeum lechleri (4) | 4 × HT41 | |
| Hordeum pubiflorum (4) | 2 × HT46, 2 × HT68 | ||
| JB027 | Hordeum lechleri (2) | 2 × HT41 | |
| Hordeum pubiflorum (1) | 1 × HT68 | ||
| JB029 | Hordeum lechleri (4) | 4 × HT41 | |
| Hordeum tetraploidum (5) | 1 × HT62, 1 × HT68, 2 × HT75, 1 × HT82 | HT68 | |
| Hordeum pubiflorum (1) | 1 × HT68 | ||
| JB030 | Hordeum parodii (4) | 4 × HT68 | |
| Hordeum lechleri (5) | 5 × HT41 | ||
| JB033 | Hordeum lechleri (5) | 4 × HT41, 1 × HT68 | |
| Hordeum pubiflorum (5) | 5 × HT46 | ||
| JB035 | Hordeum lechleri (5) | 3 × HT41, 2 × HT68 | HT68 |
| Hordeum pubiflorum (3) | 3 × HT68 | ||
| JB043 | Hordeum lechleri (5) | 5 × HT41 | |
| Hordeum tetraploidum (5) | 5 × HT46 | ||
| JB044 | Hordeum lechleri (6) | 6 × HT68 | |
| Hordeum patagonicum (5) | 5 × HT71 | ||
| Hordeum pubiflorum (4) | 4 × HT46 | ||
| JB045 | Hordeum lechleri (7) | 2 × HT41, 2 × HT62, 2 × HT68 | HT68 |
| Hordeum patagonicum (3) | 1 × HT68, 2 × HT74 | ||
| JB048 | Hordeum lechleri (3) | 2 × HT68, 1 × HT82 | |
| Hordeum patagonicum (5) | 2 × HT68, 3 × HT71 | HT68 | |
| Hordeum tetraploidum (5) | 3 × HT46, 2 × HT84 | HT82 | |
| Hordeum pubiflorum (2) | 2 × HT82 | ||
| JB050 | Hordeum lechleri (11) | 5 × HT41, 6 × HT68 | |
| Hordeum pubiflorum (4) | 4 × HT68 | HT46 | |
| Hordeum patagonicum (3) | 3 × HT46 | HT68 | |
| Hordeum comosum (3) | 1 × HT46, 1 × HT47, 1 × HT57 | ||
| JB051 | Hordeum lechleri (11) | 6 × HT41, 5 × HT68 | HT68 |
| Hordeum patagonicum (5) | 5 × HT68 | ||
| JB052 | Hordeum lechleri (5) | 5 × HT41 | |
| Hordeum patagonicum (5) | 3 × HT62, 2 × HT68 | ||
| JB053 | Hordeum lechleri (6) | 6 × HT41 | |
| Hordeum pubiflorum (6) | 1 × HT46, 3 × HT68, 2 × HT75 | ||
| Hordeum comosum (3) | 3 × HT57 | ||
| JB055 | Hordeum lechleri (5) | 5 × HT41 | |
| Hordeum parodii (3) | 3 × HT68 | ||
| Hordeum patagonicum (3) | 3 × HT64 | ||
| JB059 | Hordeum lechleri (2) | 2 × HT41 | |
| Hordeum pubiflorum (4) | 4 × HT77 | ||
| JB060 | Hordeum pubiflorum (3) | 3 × HT68 | HT68 |
| Hordeum parodii (1) | 1 × HT68 | ||
| JB236 | Hordeum comosum (2) | 2 × HT47 | |
| Hordeum pubiflorum (3) | 3 × HT85 | ||
| JB238 | Hordeum comosum (1) | 1 × HT57 | |
| Hordeum pubiflorum (3) | 1 × HT46, 1 × HT75, 1 × HT85 | ||
| JB239 | Hordeum pubiflorum (5) | 3 × HT46, 3 × HT54 | |
| Hordeum comosum (1) | 1 × HT60 | ||
| JB255 | Hordeum cordobense (1) | 1 × HT66 | |
| Hordeum stenostachys (2) | 2 × HT49 | ||
| JB257 | Hordeum cordobense (1) | 1 × HT66 | |
| Hordeum stenostachys (2) | 2 × HT51 |
Collection Site . | Species (N) . | Haplotype . | Shared Haplotype . |
|---|---|---|---|
| JB013 | Hordeum pubiflorum (5) | 5 × HT68 | |
| Hordeum patagonicum (7) | 2 × HT54, 1 × HT57, 4 × HT68 | HT68 | |
| Hordeum comosum (3) | 3 × HT46 | ||
| JB014 | Hordeum lechleri (1) | 1 × HT41 | |
| Hordeum parodii (3) | 3 × HT68 | HT57 | |
| Hordeum patagonicum (3) | 3 × HT57 | ||
| Hordeum comosum (3) | 3 × HT57 | ||
| JB015 | Hordeum lechleri (4) | 4 × HT41 | |
| Hordeum pubiflorum (2) | 1 × HT68, 1 × HT85 | HT68 | |
| Hordeum patagonicum (3) | 3 × HT85 | HT85 | |
| Hordeum comosum (3) | 3 × HT57 | ||
| Hordeum parodii (2) | 2 × HT68 | ||
| JB016 | Hordeum lechleri (3) | 3 × HT41 | |
| Hordeum pubiflorum (2) | 2 × HT68 | ||
| JB017 | Hordeum lechleri (4) | 4 × HT41 | |
| Hordeum pubiflorum (6) | 6 × HT68 | ||
| JB019 | Hordeum parodii (5) | 5 × HT46 | |
| Hordeum lechleri (5) | 5 × HT41 | ||
| JB022 | Hordeum pubiflorum (4) | 4 × HT68 | |
| Hordeum comosum (8) | 8 × HT46 | ||
| Hordeum lechleri (3) | 3 × HT41 | ||
| JB025 | Hordeum lechleri (4) | 4 × HT41 | |
| Hordeum pubiflorum (4) | 2 × HT46, 2 × HT68 | ||
| JB027 | Hordeum lechleri (2) | 2 × HT41 | |
| Hordeum pubiflorum (1) | 1 × HT68 | ||
| JB029 | Hordeum lechleri (4) | 4 × HT41 | |
| Hordeum tetraploidum (5) | 1 × HT62, 1 × HT68, 2 × HT75, 1 × HT82 | HT68 | |
| Hordeum pubiflorum (1) | 1 × HT68 | ||
| JB030 | Hordeum parodii (4) | 4 × HT68 | |
| Hordeum lechleri (5) | 5 × HT41 | ||
| JB033 | Hordeum lechleri (5) | 4 × HT41, 1 × HT68 | |
| Hordeum pubiflorum (5) | 5 × HT46 | ||
| JB035 | Hordeum lechleri (5) | 3 × HT41, 2 × HT68 | HT68 |
| Hordeum pubiflorum (3) | 3 × HT68 | ||
| JB043 | Hordeum lechleri (5) | 5 × HT41 | |
| Hordeum tetraploidum (5) | 5 × HT46 | ||
| JB044 | Hordeum lechleri (6) | 6 × HT68 | |
| Hordeum patagonicum (5) | 5 × HT71 | ||
| Hordeum pubiflorum (4) | 4 × HT46 | ||
| JB045 | Hordeum lechleri (7) | 2 × HT41, 2 × HT62, 2 × HT68 | HT68 |
| Hordeum patagonicum (3) | 1 × HT68, 2 × HT74 | ||
| JB048 | Hordeum lechleri (3) | 2 × HT68, 1 × HT82 | |
| Hordeum patagonicum (5) | 2 × HT68, 3 × HT71 | HT68 | |
| Hordeum tetraploidum (5) | 3 × HT46, 2 × HT84 | HT82 | |
| Hordeum pubiflorum (2) | 2 × HT82 | ||
| JB050 | Hordeum lechleri (11) | 5 × HT41, 6 × HT68 | |
| Hordeum pubiflorum (4) | 4 × HT68 | HT46 | |
| Hordeum patagonicum (3) | 3 × HT46 | HT68 | |
| Hordeum comosum (3) | 1 × HT46, 1 × HT47, 1 × HT57 | ||
| JB051 | Hordeum lechleri (11) | 6 × HT41, 5 × HT68 | HT68 |
| Hordeum patagonicum (5) | 5 × HT68 | ||
| JB052 | Hordeum lechleri (5) | 5 × HT41 | |
| Hordeum patagonicum (5) | 3 × HT62, 2 × HT68 | ||
| JB053 | Hordeum lechleri (6) | 6 × HT41 | |
| Hordeum pubiflorum (6) | 1 × HT46, 3 × HT68, 2 × HT75 | ||
| Hordeum comosum (3) | 3 × HT57 | ||
| JB055 | Hordeum lechleri (5) | 5 × HT41 | |
| Hordeum parodii (3) | 3 × HT68 | ||
| Hordeum patagonicum (3) | 3 × HT64 | ||
| JB059 | Hordeum lechleri (2) | 2 × HT41 | |
| Hordeum pubiflorum (4) | 4 × HT77 | ||
| JB060 | Hordeum pubiflorum (3) | 3 × HT68 | HT68 |
| Hordeum parodii (1) | 1 × HT68 | ||
| JB236 | Hordeum comosum (2) | 2 × HT47 | |
| Hordeum pubiflorum (3) | 3 × HT85 | ||
| JB238 | Hordeum comosum (1) | 1 × HT57 | |
| Hordeum pubiflorum (3) | 1 × HT46, 1 × HT75, 1 × HT85 | ||
| JB239 | Hordeum pubiflorum (5) | 3 × HT46, 3 × HT54 | |
| Hordeum comosum (1) | 1 × HT60 | ||
| JB255 | Hordeum cordobense (1) | 1 × HT66 | |
| Hordeum stenostachys (2) | 2 × HT49 | ||
| JB257 | Hordeum cordobense (1) | 1 × HT66 | |
| Hordeum stenostachys (2) | 2 × HT51 |
NOTE.—Boldface type is used to indicate diploid species in the column Species, shared haplotypes in the column Haplotype, and shared haplotypes among diploid species in the column Shared haplotype. Collection sites are detailed in table S1 (Supplementary Material online).
South American Collection Sites with Sympatric Occurrence of Two or More Hordeum Species and Chloroplast Haplotype Distribution among the Taxa
Collection Site . | Species (N) . | Haplotype . | Shared Haplotype . |
|---|---|---|---|
| JB013 | Hordeum pubiflorum (5) | 5 × HT68 | |
| Hordeum patagonicum (7) | 2 × HT54, 1 × HT57, 4 × HT68 | HT68 | |
| Hordeum comosum (3) | 3 × HT46 | ||
| JB014 | Hordeum lechleri (1) | 1 × HT41 | |
| Hordeum parodii (3) | 3 × HT68 | HT57 | |
| Hordeum patagonicum (3) | 3 × HT57 | ||
| Hordeum comosum (3) | 3 × HT57 | ||
| JB015 | Hordeum lechleri (4) | 4 × HT41 | |
| Hordeum pubiflorum (2) | 1 × HT68, 1 × HT85 | HT68 | |
| Hordeum patagonicum (3) | 3 × HT85 | HT85 | |
| Hordeum comosum (3) | 3 × HT57 | ||
| Hordeum parodii (2) | 2 × HT68 | ||
| JB016 | Hordeum lechleri (3) | 3 × HT41 | |
| Hordeum pubiflorum (2) | 2 × HT68 | ||
| JB017 | Hordeum lechleri (4) | 4 × HT41 | |
| Hordeum pubiflorum (6) | 6 × HT68 | ||
| JB019 | Hordeum parodii (5) | 5 × HT46 | |
| Hordeum lechleri (5) | 5 × HT41 | ||
| JB022 | Hordeum pubiflorum (4) | 4 × HT68 | |
| Hordeum comosum (8) | 8 × HT46 | ||
| Hordeum lechleri (3) | 3 × HT41 | ||
| JB025 | Hordeum lechleri (4) | 4 × HT41 | |
| Hordeum pubiflorum (4) | 2 × HT46, 2 × HT68 | ||
| JB027 | Hordeum lechleri (2) | 2 × HT41 | |
| Hordeum pubiflorum (1) | 1 × HT68 | ||
| JB029 | Hordeum lechleri (4) | 4 × HT41 | |
| Hordeum tetraploidum (5) | 1 × HT62, 1 × HT68, 2 × HT75, 1 × HT82 | HT68 | |
| Hordeum pubiflorum (1) | 1 × HT68 | ||
| JB030 | Hordeum parodii (4) | 4 × HT68 | |
| Hordeum lechleri (5) | 5 × HT41 | ||
| JB033 | Hordeum lechleri (5) | 4 × HT41, 1 × HT68 | |
| Hordeum pubiflorum (5) | 5 × HT46 | ||
| JB035 | Hordeum lechleri (5) | 3 × HT41, 2 × HT68 | HT68 |
| Hordeum pubiflorum (3) | 3 × HT68 | ||
| JB043 | Hordeum lechleri (5) | 5 × HT41 | |
| Hordeum tetraploidum (5) | 5 × HT46 | ||
| JB044 | Hordeum lechleri (6) | 6 × HT68 | |
| Hordeum patagonicum (5) | 5 × HT71 | ||
| Hordeum pubiflorum (4) | 4 × HT46 | ||
| JB045 | Hordeum lechleri (7) | 2 × HT41, 2 × HT62, 2 × HT68 | HT68 |
| Hordeum patagonicum (3) | 1 × HT68, 2 × HT74 | ||
| JB048 | Hordeum lechleri (3) | 2 × HT68, 1 × HT82 | |
| Hordeum patagonicum (5) | 2 × HT68, 3 × HT71 | HT68 | |
| Hordeum tetraploidum (5) | 3 × HT46, 2 × HT84 | HT82 | |
| Hordeum pubiflorum (2) | 2 × HT82 | ||
| JB050 | Hordeum lechleri (11) | 5 × HT41, 6 × HT68 | |
| Hordeum pubiflorum (4) | 4 × HT68 | HT46 | |
| Hordeum patagonicum (3) | 3 × HT46 | HT68 | |
| Hordeum comosum (3) | 1 × HT46, 1 × HT47, 1 × HT57 | ||
| JB051 | Hordeum lechleri (11) | 6 × HT41, 5 × HT68 | HT68 |
| Hordeum patagonicum (5) | 5 × HT68 | ||
| JB052 | Hordeum lechleri (5) | 5 × HT41 | |
| Hordeum patagonicum (5) | 3 × HT62, 2 × HT68 | ||
| JB053 | Hordeum lechleri (6) | 6 × HT41 | |
| Hordeum pubiflorum (6) | 1 × HT46, 3 × HT68, 2 × HT75 | ||
| Hordeum comosum (3) | 3 × HT57 | ||
| JB055 | Hordeum lechleri (5) | 5 × HT41 | |
| Hordeum parodii (3) | 3 × HT68 | ||
| Hordeum patagonicum (3) | 3 × HT64 | ||
| JB059 | Hordeum lechleri (2) | 2 × HT41 | |
| Hordeum pubiflorum (4) | 4 × HT77 | ||
| JB060 | Hordeum pubiflorum (3) | 3 × HT68 | HT68 |
| Hordeum parodii (1) | 1 × HT68 | ||
| JB236 | Hordeum comosum (2) | 2 × HT47 | |
| Hordeum pubiflorum (3) | 3 × HT85 | ||
| JB238 | Hordeum comosum (1) | 1 × HT57 | |
| Hordeum pubiflorum (3) | 1 × HT46, 1 × HT75, 1 × HT85 | ||
| JB239 | Hordeum pubiflorum (5) | 3 × HT46, 3 × HT54 | |
| Hordeum comosum (1) | 1 × HT60 | ||
| JB255 | Hordeum cordobense (1) | 1 × HT66 | |
| Hordeum stenostachys (2) | 2 × HT49 | ||
| JB257 | Hordeum cordobense (1) | 1 × HT66 | |
| Hordeum stenostachys (2) | 2 × HT51 |
Collection Site . | Species (N) . | Haplotype . | Shared Haplotype . |
|---|---|---|---|
| JB013 | Hordeum pubiflorum (5) | 5 × HT68 | |
| Hordeum patagonicum (7) | 2 × HT54, 1 × HT57, 4 × HT68 | HT68 | |
| Hordeum comosum (3) | 3 × HT46 | ||
| JB014 | Hordeum lechleri (1) | 1 × HT41 | |
| Hordeum parodii (3) | 3 × HT68 | HT57 | |
| Hordeum patagonicum (3) | 3 × HT57 | ||
| Hordeum comosum (3) | 3 × HT57 | ||
| JB015 | Hordeum lechleri (4) | 4 × HT41 | |
| Hordeum pubiflorum (2) | 1 × HT68, 1 × HT85 | HT68 | |
| Hordeum patagonicum (3) | 3 × HT85 | HT85 | |
| Hordeum comosum (3) | 3 × HT57 | ||
| Hordeum parodii (2) | 2 × HT68 | ||
| JB016 | Hordeum lechleri (3) | 3 × HT41 | |
| Hordeum pubiflorum (2) | 2 × HT68 | ||
| JB017 | Hordeum lechleri (4) | 4 × HT41 | |
| Hordeum pubiflorum (6) | 6 × HT68 | ||
| JB019 | Hordeum parodii (5) | 5 × HT46 | |
| Hordeum lechleri (5) | 5 × HT41 | ||
| JB022 | Hordeum pubiflorum (4) | 4 × HT68 | |
| Hordeum comosum (8) | 8 × HT46 | ||
| Hordeum lechleri (3) | 3 × HT41 | ||
| JB025 | Hordeum lechleri (4) | 4 × HT41 | |
| Hordeum pubiflorum (4) | 2 × HT46, 2 × HT68 | ||
| JB027 | Hordeum lechleri (2) | 2 × HT41 | |
| Hordeum pubiflorum (1) | 1 × HT68 | ||
| JB029 | Hordeum lechleri (4) | 4 × HT41 | |
| Hordeum tetraploidum (5) | 1 × HT62, 1 × HT68, 2 × HT75, 1 × HT82 | HT68 | |
| Hordeum pubiflorum (1) | 1 × HT68 | ||
| JB030 | Hordeum parodii (4) | 4 × HT68 | |
| Hordeum lechleri (5) | 5 × HT41 | ||
| JB033 | Hordeum lechleri (5) | 4 × HT41, 1 × HT68 | |
| Hordeum pubiflorum (5) | 5 × HT46 | ||
| JB035 | Hordeum lechleri (5) | 3 × HT41, 2 × HT68 | HT68 |
| Hordeum pubiflorum (3) | 3 × HT68 | ||
| JB043 | Hordeum lechleri (5) | 5 × HT41 | |
| Hordeum tetraploidum (5) | 5 × HT46 | ||
| JB044 | Hordeum lechleri (6) | 6 × HT68 | |
| Hordeum patagonicum (5) | 5 × HT71 | ||
| Hordeum pubiflorum (4) | 4 × HT46 | ||
| JB045 | Hordeum lechleri (7) | 2 × HT41, 2 × HT62, 2 × HT68 | HT68 |
| Hordeum patagonicum (3) | 1 × HT68, 2 × HT74 | ||
| JB048 | Hordeum lechleri (3) | 2 × HT68, 1 × HT82 | |
| Hordeum patagonicum (5) | 2 × HT68, 3 × HT71 | HT68 | |
| Hordeum tetraploidum (5) | 3 × HT46, 2 × HT84 | HT82 | |
| Hordeum pubiflorum (2) | 2 × HT82 | ||
| JB050 | Hordeum lechleri (11) | 5 × HT41, 6 × HT68 | |
| Hordeum pubiflorum (4) | 4 × HT68 | HT46 | |
| Hordeum patagonicum (3) | 3 × HT46 | HT68 | |
| Hordeum comosum (3) | 1 × HT46, 1 × HT47, 1 × HT57 | ||
| JB051 | Hordeum lechleri (11) | 6 × HT41, 5 × HT68 | HT68 |
| Hordeum patagonicum (5) | 5 × HT68 | ||
| JB052 | Hordeum lechleri (5) | 5 × HT41 | |
| Hordeum patagonicum (5) | 3 × HT62, 2 × HT68 | ||
| JB053 | Hordeum lechleri (6) | 6 × HT41 | |
| Hordeum pubiflorum (6) | 1 × HT46, 3 × HT68, 2 × HT75 | ||
| Hordeum comosum (3) | 3 × HT57 | ||
| JB055 | Hordeum lechleri (5) | 5 × HT41 | |
| Hordeum parodii (3) | 3 × HT68 | ||
| Hordeum patagonicum (3) | 3 × HT64 | ||
| JB059 | Hordeum lechleri (2) | 2 × HT41 | |
| Hordeum pubiflorum (4) | 4 × HT77 | ||
| JB060 | Hordeum pubiflorum (3) | 3 × HT68 | HT68 |
| Hordeum parodii (1) | 1 × HT68 | ||
| JB236 | Hordeum comosum (2) | 2 × HT47 | |
| Hordeum pubiflorum (3) | 3 × HT85 | ||
| JB238 | Hordeum comosum (1) | 1 × HT57 | |
| Hordeum pubiflorum (3) | 1 × HT46, 1 × HT75, 1 × HT85 | ||
| JB239 | Hordeum pubiflorum (5) | 3 × HT46, 3 × HT54 | |
| Hordeum comosum (1) | 1 × HT60 | ||
| JB255 | Hordeum cordobense (1) | 1 × HT66 | |
| Hordeum stenostachys (2) | 2 × HT49 | ||
| JB257 | Hordeum cordobense (1) | 1 × HT66 | |
| Hordeum stenostachys (2) | 2 × HT51 |
NOTE.—Boldface type is used to indicate diploid species in the column Species, shared haplotypes in the column Haplotype, and shared haplotypes among diploid species in the column Shared haplotype. Collection sites are detailed in table S1 (Supplementary Material online).
A further characteristic of the New World part of the network is the relatively rare occurrence of missing intermediates. This might partly be attributed to differences in sample design as we surveyed population variation particularly in Patagonia and in H. marinum in Europe and the Mediterranean. However, differences in the frequency of missing haplotypes also occur between these thoroughly sampled groups (4 vs. 14). Thus, we can exclude sampling artifacts as the sole reason for the frequency differences between the Old and the New World. In contrast to the far-reaching loss of chloroplast lineages in the Old World and particularly in the Mediterranean, the low number of missing intermediates in South America is compatible with a constantly growing effective population size, resulting in the fixation of nearly all newly arising chloroplast types (Avise 2000) and the maintenance of shared ancient polymorphisms.
Speciation Mechanisms
The occurrence of shared ancient polymorphisms among species calls for a certain number of individuals involved in a speciation event. Thus, far-reaching persistence of ancient polymorphisms indicates the subdivision of relatively large populations during speciation. In contrast, speciation due to founder events or small peripheral isolates, involving one or few individuals, should also leave a footprint in chloroplast distribution patterns (Coyne and Orr 2004). Depending on the time since speciation, we would in this case expect just one chloroplast type or only some species-specific haplotypes with shallow coalescence, all descending from the single initial progenitor of the founder individual.
The chloroplast pattern in South America with several haplotypes shared among a multitude of species indicates either repeated population subdivision or prolonged times of gene flow between formerly isolated species when they came secondarily into contact. As hybridization is insufficient to explain the chloroplast patterns found in South America (see before), we assume repeated population subdivisions involved in species formation. In contrast, the chloroplast patterns within the 2 North American species Hordeum intercedens and Hordeum pusillum, which originated from South American progenitors after 2 independent long-distance dispersals (Blattner 2006), concur with the proposed founder events as only 1 (in H. intercedens) or 2 (in H. pusillum and its hexaploid offspring Hordeum arizonicum) chloroplast types were found (fig. 2).
Biogeographical Implications
Hordeum originated about 12 Mya in Southwest Asia or adjacent areas from where it colonized eastern Asia, the New World, and South Africa. Blattner (2006) inferred at least 6 intercontinental exchanges of Hordeum lineages, involving transcontinental migrations (via Beringia) and long-distance dispersals between the Americas and between Europe and South Africa. The distribution of the chloroplast haplotypes analyzed here supports this biogeographic scenario. Long-distance dispersal from Europe to South Africa is indicated by HT16, which is shared between H. marinum subsp. marinum and its tetraploid descendant H. capense. The colonization of South America by long-distance dispersal from western North America is supported by the single chloroplast lineage (HT1) that arrived on this subcontinent. The occurrence of South American haplotypes (HT47 and 63) in the North American species H. pusillum and H. intercedens supports their South American ancestry (Blattner 2006). An entirely different pattern is shown for exchanges between Asia and North America. Three different chloroplast lineages connect both continents, converging in HT1 and 24 (fig. 2). They indicate that at least 2 extant chloroplast lineages crossed Beringia from Asia to North America and 1 (HT32 of H. jubatum) in the opposite direction. This, together with the current occurrence of H. jubatum and H. brachyantherum on both sides of the Bering Strait, indicates that Beringia was not a severe barrier to Hordeum populations during the last 4 Myr (Blattner 2006), although it was repeatedly closed as a migration route during Pleistocene glaciations.
The occurrence of the North American HT41 in tetraploid H. jubatum and hexaploid Patagonian H. lechleri, together with the high morphological similarity of both species, indicates H. jubatum as maternal progenitor of H. lechleri. This makes an intercontinental exchange from North to South America necessary to introduce the North American progenitor of H. lechleri (carrying HT41) into South America. This long-distance colonization was not postulated by Blattner (2006) on the basis of nuclear data. It emphasizes again the role of bird migration for seed exchange between Patagonia and North America. As several bird species breed in the high Arctic of North America but winter in the steppe and pampas of northern Patagonia, they provide regular transportation opportunities for seeds between both subcontinents.
Conclusions
We provide here the first chloroplast genealogy of an entire genus. It revealed that incomplete lineage sorting affects phylogenetic analysis of chloroplasts in the entire New World clade of Hordeum. The occurrence of shared haplotypes between North and South American Hordeum species means that they persisted for at least 3–4 Myr, which is the divergence time of these groups (Blattner 2006). This value is an order of magnitude higher than in, for example, Mediterranean Senecio (Asteraceae) where minimum survival times of shared chloroplast haplotypes of 0.44 Myr were estimated (Comes and Abbott 2001). Four million years is thought to be a high age, given that chloroplasts have only half the effective population sizes of nuclear inherited alleles (Avise 2000). The mechanisms behind it are not completely clear, but 2 processes should result in exceptionally large coalescence times. 1) A steady increase of individual number and, thus, effective population size will result in low lineage sorting and hence a high chance of fixation and persistence for all haplotypes (Avise 2000). 2) Isolated populations within a species buffer against lineage extinction and thus result in elongated coalescence times compared with single populations of comparable size (Nei and Takahata 1993). Both mechanisms are likely within Hordeum. Repeated dispersals of the prevalent chloroplast haplotypes to new geographic areas (Blattner 2006) eluded these haplotypes from lineage sorting in the already established populations and allowed a restart of the sorting process. In the New World, we also assume growing effective population size after the initial arrival. A restart also occurred after each onset of reproductive isolation via speciation or due to the introduction of a chloroplast type into a newly formed polyploid. Thus, the recently radiating species of the New World group of Hordeum fulfill all prerequisites for deep coalescence. However, as long survival times of chloroplast haplotypes do not seem to be restricted to Hordeum (below), expanded sampling of chloroplast sequences in other plant groups might well result in the realization that ancient polymorphisms and, therefore, old chloroplast haplotypes are not uncommon. In Hordeum, the long-term survival of chloroplast haplotypes in specific geographic areas and phylogenetic groups and their extinction in others severely influence phylogenetic inference from chloroplasts back to the deepest splits (9–12 Mya) in the genus. They resulted in pronounced differences of phylogenetic trees derived from chloroplast and nuclear loci.
A brief survey of recently published chloroplast phylogenies for zero-length branches as indicators for persisting chloroplast types revealed that these are not restricted to Hordeum. In about 90% of the studies where branch lengths were reported, indications for persisting ancient chloroplast haplotypes were found. Partly, they also occur at deep nodes in the phylogeny of genera. Ancient chloroplast haplotypes thus seem to be neither restricted to Hordeum nor uncommon. We assume therefore that in the light of our results in Hordeum, genealogical methods involving broad sampling are necessary for thorough interpretation of interspecific chloroplast data, at least if young species groups are involved.
Moreover, our data have implications not only for phylogenetic but also for phylogeographic analyses. 1) Restricting a phylogeographic analysis of chloroplast haplotypes to a single New World Hordeum species would often result in wrong interpretation of old versus young haplotypes as the oldest chloroplasts are not always in a central position or the most frequent ones (Crandall and Templeton 1993) within specific species (fig. 2). 2) Shared chloroplast types among multiple species might reflect geographical patterns, which could be older than the single species under study. Accordingly, phylogeographic analyses without knowledge of the broader chloroplast genealogy can result in the postulation of, for example, inverse colonization directions and would, thus, infer wrong species histories.
William Martin, Associate Editor
We thank Petra Oswald and Birgit Wohloier for technical assistance; Mirta Arriaga, Paula Cichero, and Roberto Gomez Cadret for kindly providing help in the organization of field work in Patagonia; Pilar Hernandez, Takao Komatsuda, the Nordic Gene Bank, and many friends and colleagues for plant material; and Peter Comes and Elizabeth Kellogg for comments on the manuscript. Financial support of the Deutsche Forschungsgemeinschaft within priority program SPP 1127 is acknowledged.
References
Albach DC, Martinez-Ortega MM, Chase MW.
Avise JC.
Bakker FT, Hellbrugge D, Culham A, Gibby M.
Baldwin BG, Sanderson MJ.
Baum BR, Bailey LG.
Blattner FR.
Blattner FR.
Bleeker W, Weber-Sparenberg C, Hurka H.
Bothmer R von, Jacobsen N, Baden C, Jørgensen RB, Linde-Laursen I.
Clement M, Posada D, Crandall KA.
Comes HP, Abbott RJ.
Comes HP, Kadereit JW.
Crandall KA, Templeton AR.
Doebley J, Bothmer R von, Larson S.
Fujii N, Ueda K, Watano Y, Shimizu T.
Hennig W.
Hudson RR.
Jakob SS, Meister A, Blattner FR.
Koch MA, Kiefer C, Ehrich D, Vogel J, Brochmann C, Mummenhoff K.
Komatsuda T, Salomon B, Bryngelsson T, Bothmer R von.
Komatsuda T, Tanno K, Salomon B, Bryngelsson T, Bothmer R von.
Linder CR, Rieseberg LH.
Mason-Gamer RJ, Holsinger KE, Jansen RK.
Nei M, Takahata N.
Nishikawa T, Salomon B, Komatsuda T, Bothmer R von, Kasowaki K.
Petersen G, Seberg O.
Petersen G, Seberg O.
Posada D, Crandall KA.
Posada D, Crandall KA.
Provan J, Russell JR, Booth A, Powell W.
Rieseberg LH, Soltis DE.
Rieseberg LH, Whitton J, Linder CR.
Sanderson MJ.
Schneider S, Roessli D, Excoffier L.
Seberg O, Frederiksen S.
Swofford DL.
Swofford DL, Olsen GJ, Waddell PJ, Hillis DM.
Taberlet P, Gielly L, Pautou G, Bouvet J.
Wendel JF, Doyle JJ.